CASCADE-ChemicAl Shift CAlculator using DEep learning
CASCADE is a stereochemically-aware online calculator for NMR chemical shifts using a graph network developed
in the Paton Group at Colorado State University.
Molecular input can be specified as SMILES or through the graphical interface.
An automated workflow executes 3D structure embedding and Merck Molecular Force Field (MMFF94) based conformer searching.
The full ensemble of optimized conformations are passed to a trained graph neural network to predict the NMR chemical shift (in ppm) for each relevant atom (C or H).
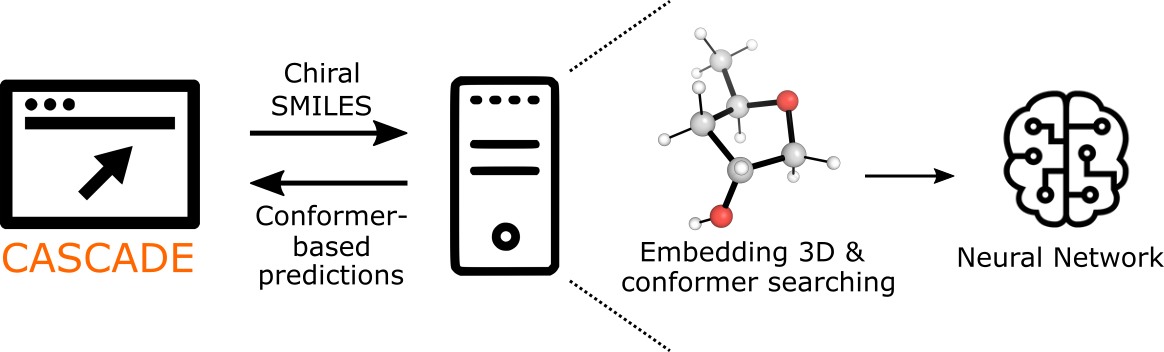
The graph neural network considers each atom as a node in a molecular graph that permits communication between atoms through edges connecting atom pairs.
Each molecule is represented by a 3D molecular graph, in which each pair of atoms are connected by a distance (i.e., as opposed to solely representing bonds in a 2D molecular graph).
CASCADE-2.0 uses the PAiNN architecture, more deatils about which can be found in the PAiNN paper and CASCADE-2.0 paper. The chemical shift predictions
are also supplemented by a confidence metric, which is indicative of the model's confidence in the prediction based on training data it has seen. Low-confidence predictions
are generally associated with
fewer training examples. For such examples, we encourage the end-user to use the neighbors feature to see examples of the most similar structures present within the training data.
Performance Evaluation
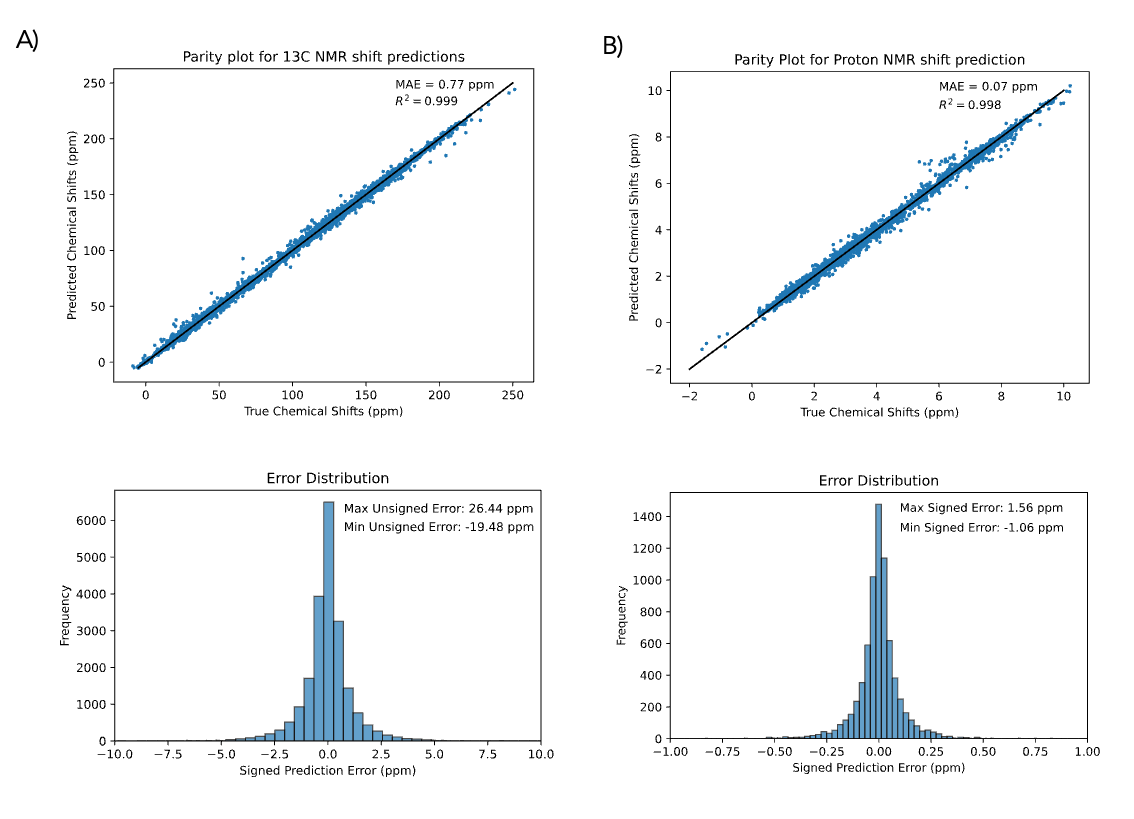
The performance of the models were evaluated against held-out test sets. The 13C NMR model was trained on a dataset of about 170,000 experimental 13C NMR shifts and tested on an external set of about 20,000 experimental 13C NMR shifts. The 1H NMR model is trained using the DFT8K dataset and tested using a set of 500 molecules with DFT 1H NMR shifts computed at the mPW1PW91/6-311+G(d,p) level of theory. The performance of both models is shown below.
Some Guidelines for using CASCADE:
- Remember to specify stereochemistry where appropriate
- The current scope of the model is neutral organic molecules consisting of C, H, N, O, S, P, F, and Cl atoms. Additionally, molecules with Si, Br, and I atoms and formal charge are supported for 13C NMR prediction.
- CASCADE generates conformers and performs energy minimizations with MMFF before predicting the chemical shifts. While there is no theoretical size-limit to this approach, extremely large molecules cause difficulties for webserver performance, particularly if multiple queries are submitted. For this reason we ask that you limit your molecules to 50 heavy atoms.
- For support with larger molecules, please install and run CASCADE on your own machine. The code is openly available on Github.
- For futher support please contact us by email: patonlab@colostate.edu
For more details, please see our paper: Real-time Prediction of 1H and 13C Chemical Shifts with DFT accuracy using a 3D Graph Neural Network. Guan, Y.; Sowndarya, S. S. V.; Gallegos, L. C.; St. John, P. C.; Paton, R. S. Chem. Sci. 2021, 23, 12012-12026. DOI:https://doi.org/10.1039/D1SC03343C.